Labs
Ausubel Lab
Professor Ausubel recently retired, his laboratory is closing down, and he is currently an emeritus Professor of Genetics at Harvard Medical School.
The Ausubel laboratory studied host-microbe interactions for the past 35 years, initially using Arabidopsis thaliana and then Caenorhabditis elegans as model genetic hosts to identify and study genes involved in the host innate immune response to pathogen attack.
Ausubel Lab Team
PI BIO
Frederick M. Ausubel is an emeritus Professor of Genetics at Harvard Medical School and the Ernst Winnacker Distinguished Investigator in the Department of Molecular Biology at Massachusetts General Hospital.
Dr. Ausubel received his B.S. in Chemistry from the University of Illinois in 1966 and his Ph.D. in Biology from the Massachusetts Institute of Technology in 1972. Formerly, he was Assistant and Associate Professor in the Department of Cellular and Developmental Biology at Harvard University.
Dr. Ausubel’s scientific work concerns host-microbe interactions. In the 1970s and 1980s, his laboratory worked on the molecular basis of symbiotic nitrogen fixation, the process by which legumes, in concert with a bacterial symbiont, convert atmospheric nitrogen into ammonia. Currently, the laboratory is investigating microbial pathogenesis and host defense in the plant Arabidopsis thaliana and the nematode Caenorhabditis elegans. The laboratory also adopted a genomics approach to study virulence in the opportunistic bacterial pathogen Pseudomonas aeruginosa strain PA14, which remarkably is a “multi-host” pathogen of both plants and animals. Of particular interest were those aspects of pathogenesis and the host innate immune response that have been conserved in evolution.
Dr. Ausubel was elected to the National Academy of Sciences in 1994, the American Academy of Microbiology in 2002, and the American Academy of Arts and Sciences in 2003. In addition to serving on a variety of editorial boards, Dr. Ausubel is founding editor of the widely-read Current Protocols in Molecular Biology.
Ausubel Lab Areas of Interest
Bacteria
Genomic Dissection of Bacterial Pathogens
To better elucidate mechanisms of pathogenicity, our laboratory is developing powerful, high-throughput models to study the infectious process and combining them with the power of bacterial genetics and genomics. Our lab has studied a number of microbial pathogens over the past several years, including species of Salmonella and Enterococcus, Staphylococcus aureus, Cryptococcus neoformans, and Fusarium oxysporum. However, the first human pathogen studied in the lab, and the best characterized, is the opportunistic pathogen, Pseudomonas aeruginosa.
Pseudomonas aeruginosa, a ubiquitous Gram-negative bacterium isolated from soil, water, and plants, is an opportunistic pathogen that infects cystic fibrosis patients, patients with thermal burns and patients who are immunodeficient or otherwise compromised. A number of years ago, we discovered that a clinical isolate, called PA14, could infect both plants and animals, and that the ability to cause disease in plants required virulence factors known to be important for pathogenicity in mammalian hosts, arguing for a conservation of mechanism in this process. Using PA14, we have developed a multi-host pathogenesis system in which the bacterium is used to infect nematodes and insects in addition to traditional cell cultures and murine hosts. These simpler model hosts are genetically tractable, reproduce more rapidly, and are smaller and cheaper to raise than their mammalian counterparts, allowing for large-scale, high-throughput assays of virulence that are impractical or unethical to perform using mice.
The Ausubel lab is currently developing PA14 tools to better characterize its behavior during infection. The most common laboratory strain of P. aeruginosa is PAO1, which was sequenced in 2000. However, this strain is generally less virulent than PA14 in a number of hosts. We have therefore sequenced the genome of PA14 to begin to understand the strain differences that account for its enhanced pathogenicity. In conjunction with the sequencing of PA14, we have created a genome-wide collection of defined, non-redundant mutations (http://ausubellab.mgh.harvard.edu/cgi-bin/pa14/home.cgi). Thirty thousand independent transposon insertion mutations were generated and each isolate was sequenced to determine the site of the lesion. From this master set, we then selected 5850 mutants corresponding to 75% of the total and approximately 80% of the non-essential PA14 ORFs. This non-redundant unigene library allows for rapid, near-saturated screens of PA14 in a number of assays, examining virulence or any other aspect of Pseudomonas biology. We used this non-redundant transposon mutant library to carry out a genome-wide screen for attenuation of PA14 virulence in C. elegans. We defined a functionally diverse 180 mutant set (representing 170 unique genes) necessary for normal levels of virulence that included both known and novel virulence factors. We examined the collection of genes required for normal levels of PA14 virulence with respect to occurrence in P. aeruginosa strain-specific genomic regions, location on putative and known genomic islands, and phylogenetic distribution across prokaryotes. Genes predominantly contributing to virulence in C. elegans showed no bias for either strain-specific regions of the P. aeruginosa genome nor putatively horizontally transferred genomic islands. Instead, within the collection of virulence-related PA14 genes, there was an over-representation of genes with a broad phylogenetic distribution that also occur with high frequency in many prokaryotic clades, suggesting that in aggregate the genes required for PA14 virulence in C. elegans are biased towards evolutionarily conserved genes.
Plants
During the past ten years, Ausubel lab has pioneered the isolation and characterization of Arabidopsis enhanced disease susceptibility (eds) mutants by direct screening, the first time that innate immunity had been subjected to rigorous and systematic genetic analysis in any eukaryotic organism. This work, as well as work from other labs, identified a variety of innate immune response pathways that regulate a multi-faceted immune response using salicylic acid (SA), jasmonic acid (JA), or ethylene (ET) as secondary messengers. Our primary goal, is to identify defense-related pathways that function in parallel (or that intersect with) the previously identified SA, JA, and ET pathways. Specifically, we have produced a substantial body of both published work and preliminary results concerning signaling pathways that respond to a highly conserved 22 amino acid bacterial flagellar peptide called Flg22. An important unanswered question in the plant innate immunity field is how the Flg22 pathway and other similar pathways that respond to so-called non-specific microbe associated molecular patterns (MAMPs) are related to the SA and JA/ET signaling pathways that primarily respond to pathogen-specific signals. We are currently using multiple approaches to further investigate this problem.
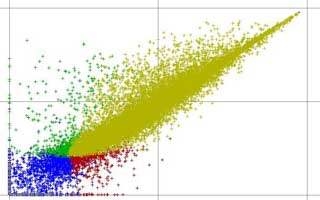
For example, we used transcriptional and metabolic profiling in Arabidopsis T-DNA mutants, coupled with the monitoring of pathogen triggered callose deposition, to identify major roles in the response to Flg22 for the plant hormone ethylene and the secondary metabolite 4-methoxy-indol-3-ylmethylglucosinolate. This study showed that the well-studied glucosinolates, previously shown to be important in avoiding damage by herbivores, are also required as a component of the plant defense response against microbial pathogens.
In another study, by monitoring transcriptional activation of GUS reporters and MAMP-elicited callose deposition, we showed that three MAMPs, Flg22, peptidoglycan, and chitin trigger a strong tissue-specific response in Arabidopsis roots, either at the elongation zone for Flg22 and peptidoglycan, or in the mature parts of the roots for chitin. Ethylene signaling, the 4-methoxy-indole-3-ylmethylglucosinolate biosynthetic pathway, and the PEN2 myrosinase, but not salicylic acid or jasmonic acid signaling, play major roles in this MAMP response. We also showed that Flg22 induces the exudation of the well-studied phytoalexin camalexin by Arabidopsis roots dependent on the cytochrome P450 CYP71A12 at physiologically significant concentrations. Interestingly, we also showed that the pathogenic bacterium Pseudomonas syringae and the beneficial microbe Pseudomonas fluorescens suppress the MAMP response in roots via the secretion of the phytotoxin coronatine, an isoleucine-jasmonic acid mimic, in the case of P. syringae, and via an unknown mechanism for P. fluorescens. The coronatine–mediated suppression of MAMP responses, including transcription of MYB51, a transcription factor involved in MAMP signaling pathways, requires the E3 ubiquitin ligase COI1 and the transcription factor JIN1/MYC2, but does not rely on salicylic acid-jasmonic acid antagonism. These latter experiments demonstrate the presence of highly orchestrated and tissue-specific MAMP responses in roots and pathogen-encoded mechanisms to block these MAMP-elicited signaling pathways.

A final Arabidopsis project concerns the underlying mechanisms by which P. aeruginosa and P. syringae are able to infect Arabidopsis leaves. In the case of P. aeruginosa, we have found that the disaccharide trehalose is a key P. aeruginosa virulence factor. We are currently testing several hypotheses concerning the role of trehalose in the infection process, including the extraction of nutrients from plant cells. Interestingly, the ability to extract nutrients from mesophyll cells is also a key factor underlying P. syringae virulence. Conversely, we have discovered that the withdrawal of nutrients from the intercellular spaces in the leaf, especially amino acids, is an important Arabidopsis defense response to pathogen attack.
Worms
The Ausubel lab uses genetic and whole genome approaches to investigate the molecular basis of microbial pathogenesis and host innate immunity.
The vertebrate innate immune system serves as the first line of defense against microbial invaders, and activates the other branch of immunity, the adaptive immune system. Although the adaptive immune system, involving the clonal selection of lymphocytes, is unique to vertebrates, the innate immune response has ancient origins. Innate immunity is characterized by receptors that are activated by signals indicative of pathogens, often referred to as microbe-associated molecular patterns (MAMPs). MAMPs include conserved microbial cell wall molecules such as lipopolysaccharide and peptidoglycan. MAMP receptors activate downstream signaling cascades; ultimately, a set of immune effectors is activated.
Comprehensive genetic analyses of the molecules that comprise the innate immune response have only recently been initiated. Model systems, including Drosophila melanogaster, have proven invaluable in the identification of new components of innate immunity, many of which have conserved roles in vertebrates. Numerous studies now suggest that the fundamental mechanisms of host defense against pathogen invasion are conserved across metazoans, highlighting the importance of continued genetic analysis of the immune system in model species.
Our lab has pioneered the use of the nematode Caenorhabditis elegans as a new model host to study the innate immune response to diverse pathogens. This nematode-pathogen model system is advantageous because worms are simply fed various bacteria or yeast in place of their usual food source of Escherichia coli (strain OP50) and observed for their survival over time. A remarkably large number of human pathogens have been found to be lethal to C. elegans in this assay, including the Gram-positive bacteria Staphyloccus aureus and Enterococcus faecalis, the Gram-negative bacteria Pseudomonas aeruginosa, and Salmonella enterica, and the yeasts Cryptococcus neoformans and Candida albicans.
Powerful biochemical, molecular, and forward and reverse genetic tools are readily available in C. elegans, permitting comprehensive analysis of both bacterial virulence factors and host defense factors. Both pathogen and host immune molecules have been identified that overlap significantly with known factors in mammalian models, indicating the evolutionary conservation of pathogenesis and host immune response and the relevance of the nematode model system to vertebrates. Additionally, novel bacterial genes with previously unknown roles in pathogenesis have been discovered using the C. elegans model system and subsequently shown to be required for pathogenesis in mice. Thus, the C. elegans-pathogen system has the potential for the rapid identification and characterization of the signaling molecules and complex pathways that comprise the innate immune response and the bacterial factors that function in the pathogenesis process.
Using both forward and reverse genetic approaches, we have identified several parallel immune signaing pathways, including at least six C. elegans genes (tir-1, nsy-1, sek-1, pmk-1, mek-1, and vhp-1) that function in a conserved p38 MAPK immune signaling pathway. Tir-1 encodes a TIR-domain-containing protein that functions upstream of a MAPK cascade encoded by nsy-1 (MAPKKK), sek-1 (MAPKK) and pmk-1 (p38 MAPK). NSY-1 and SEK-1 are required for PMK-1 activation and the MAPKK encoded by mek-1 enhances PMK-1 (p38) phosphorylation. Vhp-1 encodes a MAPK phosphatase and appears to work in conjunction with SEK-1 and MEK-1 to fine-tune levels of PMK-1 activation. We have also shown that long-lived mutants in the DAF-2 insulin-like signaling pathway are highly resistant to both Gram-positive and Gram-negative pathogens and that this resistance is suppressed by mutations in the p38 MAPK pathway. Our microarray studies of the genes regulated by the p38 MAPK pathway and the DAF-2-DAF-16 pathway suggest that the p38 MAPK pathway acts in parallel to DAF-2-DAF-16 pathway to regulate immune gene expression. Interestingly, C. elegans responds differentially to infection by P. aeruginosa and S. aureus. P. aeruginosa infection activates the p38 MAPK signaling pathway as well as at least two additional parallel pathways involving the G protein coupled receptor FSHR-1 or the transcription factor ZIP-2. S. aureus, on the other hand, primarily activates an immune response pathway that involves the C. elegans homolog of beta catenin.
Intestinal epithelial cells (IECs) are exposed to a complex environment populated with both innocuous and pathogenic microbes. To mount an appropriate immune response, it is essential for IECs to distinguish between pathogens and innocuous or beneficial microbes. It is mostly unknown, however, how IECs recognize virulent microbes while ignoring or promoting the growth of their commensals or non-pathogenic counterparts. It is currently believed that pathogens are primarily detected by host pattern recognition receptors (PRRs) that recognize MAMPs. This model, however, fails to address how IECs specifically identify pathogens since both pathogenic and innocuous microbes synthesize the same MAMPs. Alternatively, there is mounting evidence that host immune responses can be activated by indirect evidence of infection. Indeed, in a process called “effector-triggered immunity” it is has been well established that plants recognize pathogens by the effects of pathogen-encoded virulence factors on host signaling pathways. In metazoans indirect indicators of pathogen activity have been referred to as “patterns-of-pathogenesis” or “damage-associated molecular patterns” (DAMPs). To help elucidate the mechanisms underlying pathogen recognition in metazoans, we investigated how P. aeruginosa triggers intestinal innate immunity in C. elegans. To test the hypothesis that P. aeruginosa-encoded toxins are responsible for activating host immunity, we engineered a normally non-pathogenic E. coli strain to produce the P. aeruginosa-encoded AB toxin Exotoxin A (ToxA), which catalyzes the ADP-ribosylation of elongation factor 2 (EF2) by targeting the EF2 diphthamide moiety, a post-translationally modified histidine, thereby blocking protein synthesis. We found that ToxA synthesized in E. coli activates a strong immune response, activating a significant subset of genes normally induced when C. elegans is infected with P. aeruginosa. Interestingly, ToxA not only activates an immune response in C. elegans, but at least two major immune response pathways in C. elegans involving the ATF-7 and ZIP-2 transcription factors, protect C. elegans from ToxA-mediated lethality. We investigated the mechanism by which ToxA activates the C. elegans immune response. First, we found that enzymatically inactive ToxA protein failed to activate an immune response. Similarly, C. elegans mutants that are unable to synthesize the diphthimide residue, the target of ToxA, did not respond to ToxA. Importantly, the transcriptional inhibitor hygromycin activated an immune response that was similar to the response activated by ToxA. Thus, ToxA appears to upregulate the C. elegans immune response by disruption of host protein translation. Our data show that C. elegans has a surveillance mechanism that recognizes ToxA through its effect on protein translation rather than by direct recognition of either ToxA or ribosylated EF2.
Finally, we have taken advantage of the small size of C. elegans to develop high throughput chemical screens for compounds that function as novel anti-infectives that enhance host immunity or block pathogen virulence. For example, we carried out an automated, high throughput screen of 37,200 compounds and natural product extracts for those that enhance survival of C. elegans infected with E. faecalis. The screen used a robot to accurately dispense live, infected animals into 384-well plates, and automated microscopy and image analysis to generate quantitative, high content data. We identified 28 compounds and extracts that were not previously reported to have antimicrobial properties, including 6 structural classes that cure infected C. elegans animals but do not affect the growth of the pathogen in vitro, thus acting by a mechanism of action distinct from antibiotics currently in clinical use. Recently we have shown that one of these small molecules, RPW-24, protects C. elegans from bacterial infection by stimulating the host immune response of the nematode. Using transcriptome profiling, epistasis pathway analyses with C. elegans mutants, and an RNAi screen, we show that RPW-24 promotes resistance to Pseudomonas aeruginosa infection by inducing the transcription of a remarkably small number of C. elegans genes (~1.3% of all genes) in a manner that partially depends on the evolutionarily-conserved p38 MAP kinase pathway and the transcription factor ATF-7. These data show that the immunostimulatory activity of RPW-24 is required for its efficacy and define a novel C. elegans-based strategy to identify compounds with activity against antibiotic-resistant bacterial pathogens. Currently, we are carrying out a second high throughput screen that utilizes P. aeruginosa-infected C. elegans.
Ausubel Lab Resources
Resources
Molecular Markers
CAPS
Cleaved Amplified Polymorphic Sequences can be used to unambiguously map an Arabidopsis gene to one of the 10 Arabidopsis chromosome arms.
SNAP
Following the design of new SNAP markers using the SNAPER program, it is necessary to test the new primer sets to select for primer pairs that show the range of specificity required for their use as molecular markers.
Strain Collection
The Strain Collection Database (SCD) is designed to keep track of strain collection information in the Ausubel Lab. It provides a web interface for registered and public users. Registered users are authorized to search and maintain the database. Public users have access to information on published strain records.



