Quick Links
Worms
The Ausubel lab uses genetic and whole genome approaches to investigate the molecular basis of microbial pathogenesis and host innate immunity.
The vertebrate innate immune system serves as the first line of defense against microbial invaders, and activates the other branch of immunity, the adaptive immune system. Although the adaptive immune system, involving the clonal selection of lymphocytes, is unique to vertebrates, the innate immune response has ancient origins. Innate immunity is characterized by receptors that are activated by signals indicative of pathogens, often referred to as microbe-associated molecular patterns (MAMPs). MAMPs include conserved microbial cell wall molecules such as lipopolysaccharide and peptidoglycan. MAMP receptors activate downstream signaling cascades; ultimately, a set of immune effectors is activated.
Comprehensive genetic analyses of the molecules that comprise the innate immune response have only recently been initiated. Model systems, including Drosophila melanogaster, have proven invaluable in the identification of new components of innate immunity, many of which have conserved roles in vertebrates. Numerous studies now suggest that the fundamental mechanisms of host defense against pathogen invasion are conserved across metazoans, highlighting the importance of continued genetic analysis of the immune system in model species.
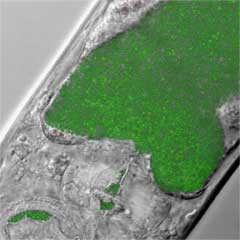
Our lab has pioneered the use of the nematode Caenorhabditis elegans as a new model host to study the innate immune response to diverse pathogens. This nematode-pathogen model system is advantageous because worms are simply fed various bacteria or yeast in place of their usual food source of Escherichia coli (strain OP50) and observed for their survival over time. A remarkably large number of human pathogens have been found to be lethal to C. elegans in this assay, including the Gram-positive bacteria Staphyloccus aureus and Enterococcus faecalis, the Gram-negative bacteria Pseudomonas aeruginosa, and Salmonella enterica, and the yeasts Cryptococcus neoformans and Candida albicans.
Powerful biochemical, molecular, and forward and reverse genetic tools are readily available in C. elegans, permitting comprehensive analysis of both bacterial virulence factors and host defense factors. Both pathogen and host immune molecules have been identified that overlap significantly with known factors in mammalian models, indicating the evolutionary conservation of pathogenesis and host immune response and the relevance of the nematode model system to vertebrates. Additionally, novel bacterial genes with previously unknown roles in pathogenesis have been discovered using the C. elegans model system and subsequently shown to be required for pathogenesis in mice. Thus, the C. elegans-pathogen system has the potential for the rapid identification and characterization of the signaling molecules and complex pathways that comprise the innate immune response and the bacterial factors that function in the pathogenesis process.
Using both forward and reverse genetic approaches, we have identified several parallel immune signaing pathways, including at least six C. elegans genes (tir-1, nsy-1, sek-1, pmk-1, mek-1, and vhp-1) that function in a conserved p38 MAPK immune signaling pathway. Tir-1 encodes a TIR-domain-containing protein that functions upstream of a MAPK cascade encoded by nsy-1 (MAPKKK), sek-1 (MAPKK) and pmk-1 (p38 MAPK). NSY-1 and SEK-1 are required for PMK-1 activation and the MAPKK encoded by mek-1 enhances PMK-1 (p38) phosphorylation. Vhp-1 encodes a MAPK phosphatase and appears to work in conjunction with SEK-1 and MEK-1 to fine-tune levels of PMK-1 activation. We have also shown that long-lived mutants in the DAF-2 insulin-like signaling pathway are highly resistant to both Gram-positive and Gram-negative pathogens and that this resistance is suppressed by mutations in the p38 MAPK pathway. Our microarray studies of the genes regulated by the p38 MAPK pathway and the DAF-2-DAF-16 pathway suggest that the p38 MAPK pathway acts in parallel to DAF-2-DAF-16 pathway to regulate immune gene expression. Interestingly, C. elegans responds differentially to infection by P. aeruginosa and S. aureus. P. aeruginosa infection activates the p38 MAPK signaling pathway as well as at least two additional parallel pathways involving the G protein coupled receptor FSHR-1 or the transcription factor ZIP-2. S. aureus, on the other hand, primarily activates an immune response pathway that involves the C. elegans homolog of beta catenin.
Intestinal epithelial cells (IECs) are exposed to a complex environment populated with both innocuous and pathogenic microbes. To mount an appropriate immune response, it is essential for IECs to distinguish between pathogens and innocuous or beneficial microbes. It is mostly unknown, however, how IECs recognize virulent microbes while ignoring or promoting the growth of their commensals or non-pathogenic counterparts. It is currently believed that pathogens are primarily detected by host pattern recognition receptors (PRRs) that recognize MAMPs. This model, however, fails to address how IECs specifically identify pathogens since both pathogenic and innocuous microbes synthesize the same MAMPs. Alternatively, there is mounting evidence that host immune responses can be activated by indirect evidence of infection. Indeed, in a process called “effector-triggered immunity” it is has been well established that plants recognize pathogens by the effects of pathogen-encoded virulence factors on host signaling pathways. In metazoans indirect indicators of pathogen activity have been referred to as “patterns-of-pathogenesis” or “damage-associated molecular patterns” (DAMPs). To help elucidate the mechanisms underlying pathogen recognition in metazoans, we investigated how P. aeruginosa triggers intestinal innate immunity in C. elegans. To test the hypothesis that P. aeruginosa-encoded toxins are responsible for activating host immunity, we engineered a normally non-pathogenic E. coli strain to produce the P. aeruginosa-encoded AB toxin Exotoxin A (ToxA), which catalyzes the ADP-ribosylation of elongation factor 2 (EF2) by targeting the EF2 diphthamide moiety, a post-translationally modified histidine, thereby blocking protein synthesis. We found that ToxA synthesized in E. coli activates a strong immune response, activating a significant subset of genes normally induced when C. elegans is infected with P. aeruginosa. Interestingly, ToxA not only activates an immune response in C. elegans, but at least two major immune response pathways in C. elegans involving the ATF-7 and ZIP-2 transcription factors, protect C. elegans from ToxA-mediated lethality. We investigated the mechanism by which ToxA activates the C. elegans immune response. First, we found that enzymatically inactive ToxA protein failed to activate an immune response. Similarly, C. elegans mutants that are unable to synthesize the diphthimide residue, the target of ToxA, did not respond to ToxA. Importantly, the transcriptional inhibitor hygromycin activated an immune response that was similar to the response activated by ToxA. Thus, ToxA appears to upregulate the C. elegans immune response by disruption of host protein translation. Our data show that C. elegans has a surveillance mechanism that recognizes ToxA through its effect on protein translation rather than by direct recognition of either ToxA or ribosylated EF2.
Finally, we have taken advantage of the small size of C. elegans to develop high throughput chemical screens for compounds that function as novel anti-infectives that enhance host immunity or block pathogen virulence. For example, we carried out an automated, high throughput screen of 37,200 compounds and natural product extracts for those that enhance survival of C. elegans infected with E. faecalis. The screen used a robot to accurately dispense live, infected animals into 384-well plates, and automated microscopy and image analysis to generate quantitative, high content data. We identified 28 compounds and extracts that were not previously reported to have antimicrobial properties, including 6 structural classes that cure infected C. elegans animals but do not affect the growth of the pathogen in vitro, thus acting by a mechanism of action distinct from antibiotics currently in clinical use. Recently we have shown that one of these small molecules, RPW-24, protects C. elegans from bacterial infection by stimulating the host immune response of the nematode. Using transcriptome profiling, epistasis pathway analyses with C. elegans mutants, and an RNAi screen, we show that RPW-24 promotes resistance to Pseudomonas aeruginosa infection by inducing the transcription of a remarkably small number of C. elegans genes (~1.3% of all genes) in a manner that partially depends on the evolutionarily-conserved p38 MAP kinase pathway and the transcription factor ATF-7. These data show that the immunostimulatory activity of RPW-24 is required for its efficacy and define a novel C. elegans-based strategy to identify compounds with activity against antibiotic-resistant bacterial pathogens. Currently, we are carrying out a second high throughput screen that utilizes P. aeruginosa-infected C. elegans.


